XB-IMG-127087
Xenbase Image ID: 127087
|
|
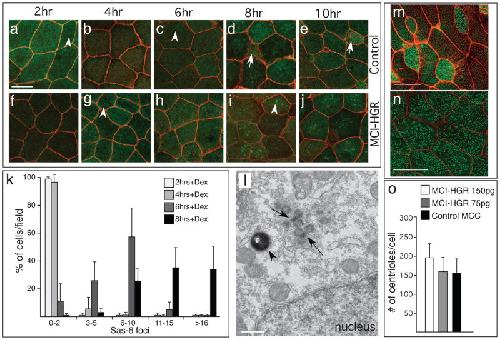
Figure 4. MCI induced centriole assembly(a–j) Shown are confocal images of the skin of control (a–e) or MCI-HGR RNA injected embryos (f–j), also expressing SAS6-GFP (green) and mRFP(red). Embryos were treated with DEX at stage 11.5, followed by fixation at the time intervals indicated. In control embryos, Sas6-GFP lightly labels centrioles (a,c arrowhead) in outer cells and basal bodies/centrioles in MCCs (d,e, arrow). By contrast, large bright Sas6-GFP foci are induced in outer cells by MCI-HGR (g,i arrowhead). Scale bar (a)=20microns (k) Plot showing the average fraction of cells (±s.d.) per field with different SAS-GFP foci number at the indicated time point, based on twelve fields taken from four embryos injected with MCI-HGR RNA. In control embryos, every outer cell scored contained 0–2 centrioles labeled with Sas6-GFP (data not shown). (l) Shown is a TEM image taken of an embryo injected with MCI-HGR RNA and treated with DEX for 8hrs. Structures similar to deuterosomes (arrows) in a pigmented (arrowhead) outer cell are located apical to the nucleus. Scale bar=500nm. (m–n) Shown are confocal images of the skin in embryos injected with mRFP and Hyls1-GFP RNA to mark cell boundaries (red) and centrioles (green), respectively, either alone (m) or with MCI-HGR RNA (n). At stage 11.5, embryos were treated with Dex, and then fixed at stage 28. Scale bars= 20microns (o) The average centriole number per cell (±s.d.) is plotted for control and MCI-HGR expressing embryos, using data obtained by scoring cells in at least three fields from five embryos. The slight increase in basal body number obtained with 150pg is significantly different (p<0.05) from control MCCs. Image published in: Stubbs JL et al. (2012) Image downloaded from an Open Access article in PubMed Central. Image reproduced on Xenbase with permission of the publisher and the copyright holder. Larger Image Printer Friendly View |
