XB-IMG-130638
Xenbase Image ID: 130638
|
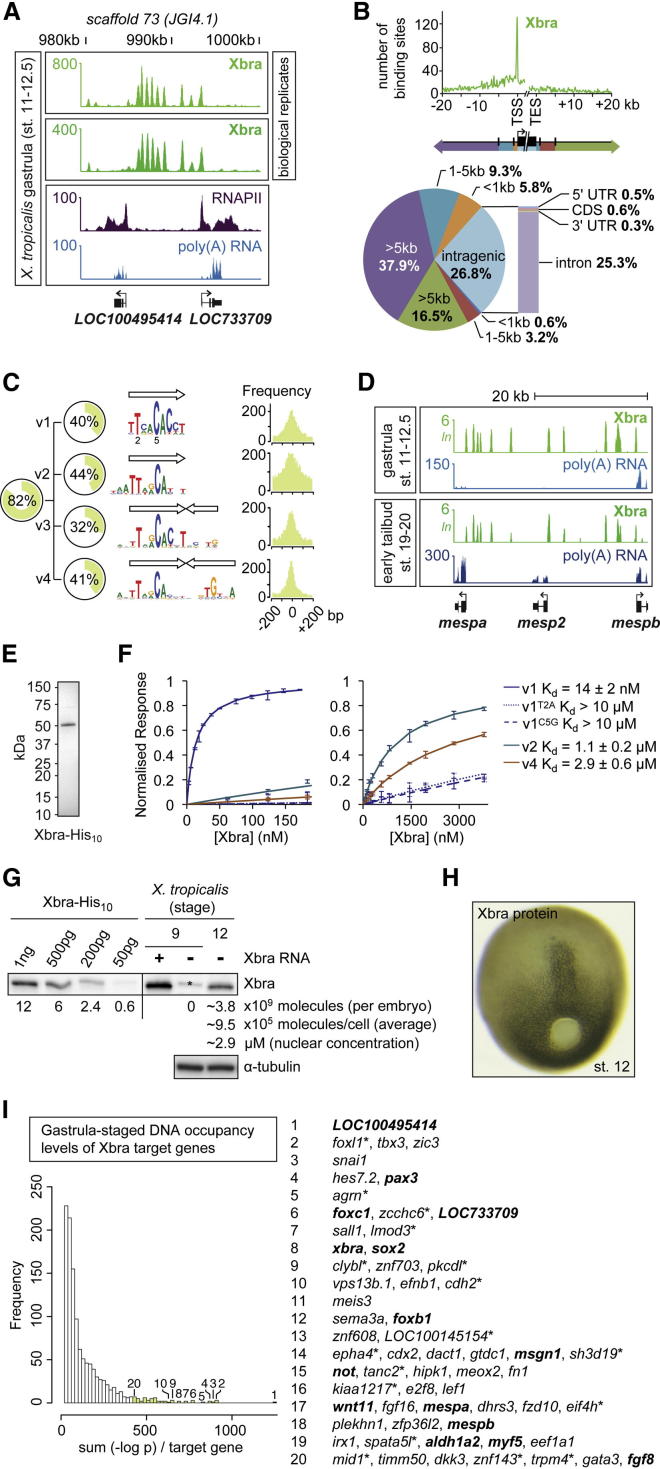
Figure 1. Xbra Is Stably Recruited to Mono- and Dimeric Motif Variants in the X. tropicalis Genome during Early Embryogenesis(A) Excerpt of normalized Xbra binding at gastrula stage. RNAPII and poly(A) RNA profile from Akkers et al. (2009).(B) Genomic distribution of Xbra binding sites (FDR ≤ 1%) relative to the start (TSS) and end (TES) of transcription of nearest target genes.(C) De novo motif discovery analysis of Xbra-bound regions with coverage, sequence logo, and positional distribution for each T-box motif variant (v1–v4). Arrow indicates monomeric binding site.(D) Comparison of Xbra binding at gastrula and early tail bud stage near mesp gene cluster with poly(A) RNA profile from Akkers et al. (2009) and this study.(E) Coomassie staining of 0.5 μg purified Xbra-His10 run on a SDS-polyacrylamide gel.(F) Surface plasmon resonance diagrams (normalized response versus Xbra concentration) including Kd values for the interaction between native Xbra protein and different DNA motifs (v1, v2, and v4). Superscript T2A and C5G refer to base changes introduced in v1.(G) Quantification of Xbra protein levels in midgastrula embryos (stage 12) by western blotting with standard curve of purified Xbra-His10 as indicated. Positive control, pregastrula embryo (stage 9) injected with RNA encoding untagged Xbra. Protein extracts equivalent to two embryos at stages 9 (negative control) and 12 were loaded. Asterisk marks nonspecific band seen at stage 9. The same band is present at the same intensity in the absence of Xbra at stage 12 (data not shown), and its intensity was therefore subtracted from the Xbra band for quantification. Further calculations (molecules/cell) and nuclear concentrations (μM) are based on an estimated 4,000 Xbra-positive cells at stage 12 ([H]; Cooke, 1979), a nuclear envelope (sphere) surface of ∼300 μm2 (Levy and Heald, 2010), and 90% of Xbra being nuclear. Loading control, α-tubulin.(H) Whole-mount immunohistochemistry of Xbra protein in a midgastrula embryo (stage 12).(I) Histogram of nearest gene-associated Xbra binding levels as detected by ChIP-seq. The asterisk indicates genes with nearest Xbra binding >10 kb from TSS. Genes in bold are mentioned elsewhere in this study.See also Figure S1 and Table S1. Image published in: Gentsch GE et al. (2013) © 2013 The Authors. Creative Commons Attribution license Larger Image Printer Friendly View |
