XB-IMG-130645
Xenbase Image ID: 130645
|
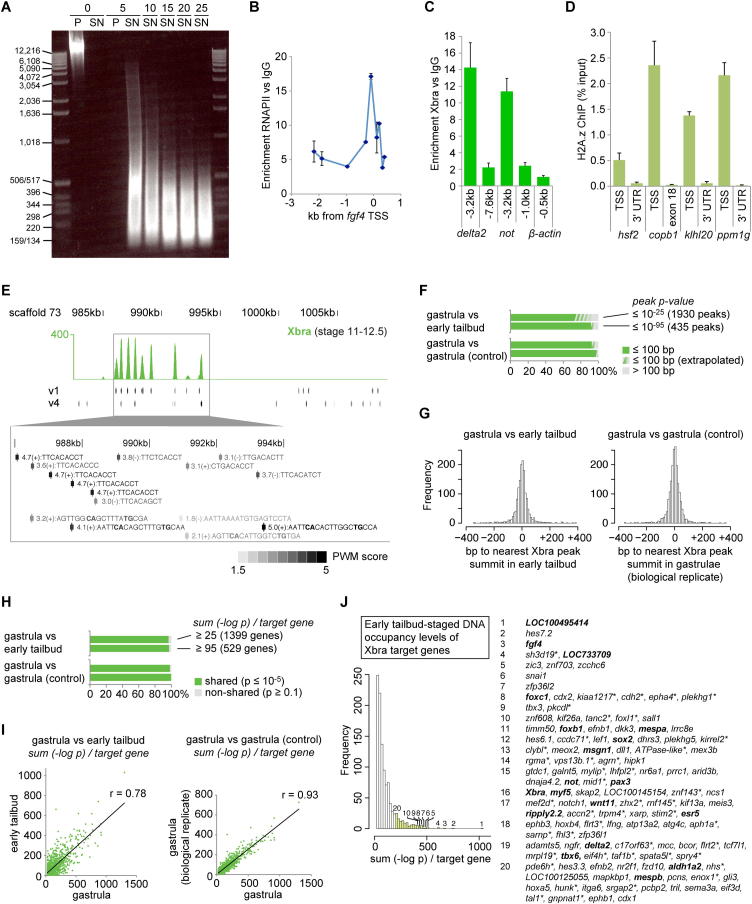
Figure S1. Development of the ChIP Protocol for Xenopus Embryos Suitable for Deep Sequencing and Near-Identical Xbra Binding Characteristics at Gastrula and Early Tail Bud Stage, Related to Figure 1(A) Increasing rounds (5, 10, 15, 20 and 25) of sonication caused cross-linked chromatin extracted from X. tropicalis gastrula embryos (stage 12) to be solubilised (compare pellet [P] to supernatant [SN] before and after 5 rounds of sonication) and sheared to fragments of the indicated sizes. Ethidium Bromide staining of genomic DNA size-separated by gel electrophoresis.(B) RNA polymerase II (RNAPII) ChIP-qPCR on X. laevis gastrula embryos (stage 12) showed RNAPII recruitment to the TSS of fgf4 (also known as eFGF(i) promoter, AF078081) (Casey et al., 1998). DNA enrichment of RNAPII versus IgG ChIP plotted against distance (kb) to transcription start site (TSS) of fgf4. Error bars, s.d. of biological duplicates.(C) Xbra ChIP-qPCR on X. tropicalis gastrula embryos (stage 12) showed Xbra binding to 3.2-kb upstream region of the delta2 and notochord (not) gene (10- to 15-fold DNA enrichment of Xbra versus IgG ChIP). not, delta2 and β-actin (negative control) loci were used as the ‘gold-standard’ controls to improve the efficiency and signal-to-noise ratio of the Xbra ChIP on X. tropicalis embryos. Positive control regions were selected based on the homology to respective target genes determined by ChIP on zebrafish mid-gastrula embryos (Morley et al., 2009) and the availability of canonical Xbra recognition sites determined in vitro (Conlon et al., 2001). Error bars, s.d. of biological replicates (n = 2-4).(D) The ChIP protocol devised for Xenopus embryos was also applicable to cross-linked zebrafish embryos. H2A.z ChIP-qPCR showed significantly more DNA occupancy (plotted as percentage of DNA input) of the histone variant H2A.z at the TSS (proximal promoter) than at the end (3′ UTR or last exon) of transcriptionally active genes (hsf2, copb1, klhl20 and ppm1g) in zebrafish bud-staged embryos. Error bars, s.d. of technical triplicates. This binding profile is in accordance to findings reported for the human genome (Barski et al., 2007).(E) Illustration of overlap between Xbra binding as shown in Figure 1A and occurrence of motif variant v1 and v4 (Figure 1C). A magnification of the intergenic region between LOC100495414 and LOC733709 shows the sequences found on the + or - strand that match the position weight matrix (PWM) of v1 and v4 with the indicated score.(F) Co-localization (%) of peak summits (distance ≤ 100 bp) between gastrula and early tailbud stage under conditions of high stringency (p ≤ 10−25 or 10−95). Extrapolation was based on visual inspection of Xbra binding profiles (see (H)). The comparison of biological replicates of gastrula-specific Xbra binding profile was used as a control in (F)-(I).(G) Histogram of distances between corresponding Xbra peaks (FDR ≤ 1%) detected in gastrula and early tailbud embryos.(H) Overlap (%) of Xbra target genes between gastrula and early tailbud stages under the condition that the sum of Xbra binding exceeds 25 or 95 [-log p] per gene at gastrula or early tailbud stage. Visual inspection of Xbra binding profiles revealed that 44 of the top 50 of non-shared target genes sorted by the sum [-log p] were incorrectly declared as the peak caller failed to identify existent peaks (p ≤ 0.1). The same inspections were carried out for non-shared target genes at tailbud stage (34/40 incorrect) and non-shared target genes of biological replicates (21/45 incorrect).(I) Linear correlation of DNA occupancy levels (sum [-log p]) at individual target genes shared between gastrula and early tailbud embryos. Deviation from this linear positive correlation is reflected by a decreasing Pearson’s correlation coefficient r.(J) Histogram (bins of 20 [-log p]) of nearest gene-associated Xbra binding levels as detected by ChIP-Seq at early tailbud stage (sum [-log p] ≥ 25). ∗ indicates genes with nearest Xbra binding further than 10 kb away from TSS. Identified target genes in bold are mentioned elsewhere in this study. Image published in: Gentsch GE et al. (2013) © 2013 The Authors. Creative Commons Attribution license Larger Image Printer Friendly View |
