XB-IMG-133538
Xenbase Image ID: 133538
|
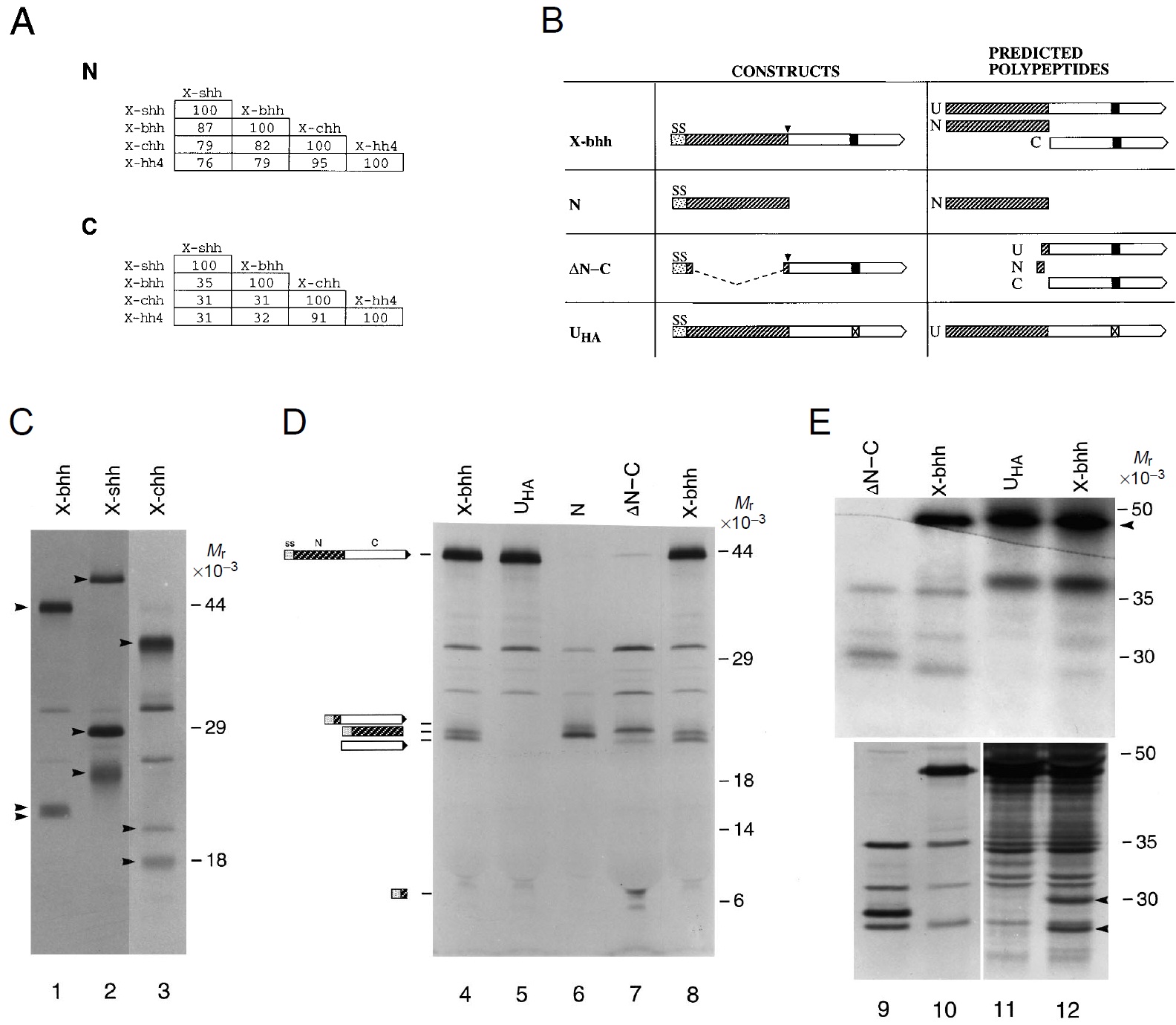
Fig. 1. Autoproteolytic cleavage of Xenopus hh proteins in vitro and
in embryos. (A) Percentage of amino acid identity between the
predicted N and C domains of the Xenopus hh gene family members
(Ekker et al., 1995). (B) Depiction of constructs encoding full-length
X-bhh, N, DN-C, and UHA. The polypeptides predicted to be formed
in vivo after translation and cleavage of the signal sequence, and the
autoproteolytic cleavage of the full-length polypeptide, are shown to
the right, and are described in Materials and methods. SS denotes the
signal sequence, U refers to the unprocessed polypeptide after
cleavage of the signal sequence, N depicts the amino-terminal region
after signal sequence cleavage and after autoproteolysis (at the site
indicated by the downward arrowhead), and C denotes the carboxyterminal
domain after autoproteolysis. The filled box in C denotes a
histidine at position 270, and the box with a check denotes mutation
of histidine 270 to an alanine. (C) Processing of X-bhh (lane 1), Xshh
(lane 2), and X-chh (lane 3) upon translation in vitro. Each lane
contains three hh-associated protein products (indicated by
arrowheads), of which the two smaller products arise by
autoproteolytic cleavage of the larger unprocessed form (see text and
D below). The two X-bhh cleavage products are of similar size, and
appear as a doublet in lane 1 (see D below). (D) Detailed analysis of
X-bhh processing in vitro. The X-bhh open reading frame was
mutated to yield UHA, N, and DN-C constructs as diagrammed in B.
Cartoons clarifying the region of X-bhh present in the translation
product are shown to the left of lane 4. Lanes 4 and 8: translation of
wild-type X-bhh. Lane 5: translation of UHA. Lane 6: translation of
N. The protein product comigrates with a fragment generated by
autoproteolysis of X-bhh (compare lane 6 with lanes 4 and 8). Lane
7: translation of DN-C whose primary translation product undergoes
autoproteolysis (refer to cartoon). The lower of the two bands within
the doublet comigrates with a fragment generated by autoproteolysis
of X-bhh (compare lanes 7 and 8 with reference to the cartoon), and
the band migrating near the 6´103 Mr marker is the small N-terminal
fragment remaining after autoproteolysis (refer to cartoon). (E)
Processing of X-bhh in embryos. X-bhh or UHA were co-injected
with [35S]methionine into embryos and the resulting extracts were
immunoprecipitated with an antibody to the carboxy region of X-bhh
(see Materials and methods). The upper gel is useful solely for
showing the presence of full-length X-bhh denoted by an arrowhead.
The lower gel was overexposed to resolve lower molecular mass
species arising by processing of X-bhh. Lane 9 (both gels): proteins
generated from in vitro translation of DN-C. Lane 10 (both gels): in
vitro translation of X-bhh. Lane 11 (both gels): Immunoprecipitation
of embryo extracts with a C-terminal antibody after injection of UHA
demonstrates the presence of full-length X-bhh polypeptide
(arrowhead, upper gel), but no bands co-migrating with C-terminal
polypeptides (lower gel). Lane 12 (both gels): Immunoprecipitation
of embryo extracts after injection of X-bhh RNA demonstrates the
presence of full length X-bhh in the upper gel. In lane 12 of the
lower gel, two lower molecular mass bands (arrowheads) are noted,
which are absent from the UHA-injected embryos (lane 11), and
absent from uninjected embryos (not shown). The lower of these two
bands comigrates with C generated by in vitro translation of X-bhh
(lane 10) or DN-C (lane 9). The approximately 30´103 Mr band in
lane 12 (arrowhead) is presumed to be a modification of the C
protein, possibly glycosylation at a predicted N-linked glycosylation
site (Ekker et al., 1995). Unmarked bands are not hh-derived as
determined by immunoprecipitation of labeled embryos not injected
with X-bhh RNAs (not shown). Image published in: Lai CJ et al. (1995) Copyright © 1995. Image reproduced with permission of the publisher and the copyright holder. This is an Open Access article distributed under the terms of the Creative Commons Attribution License. Larger Image Printer Friendly View |
