XB-IMG-137995
Xenbase Image ID: 137995
|
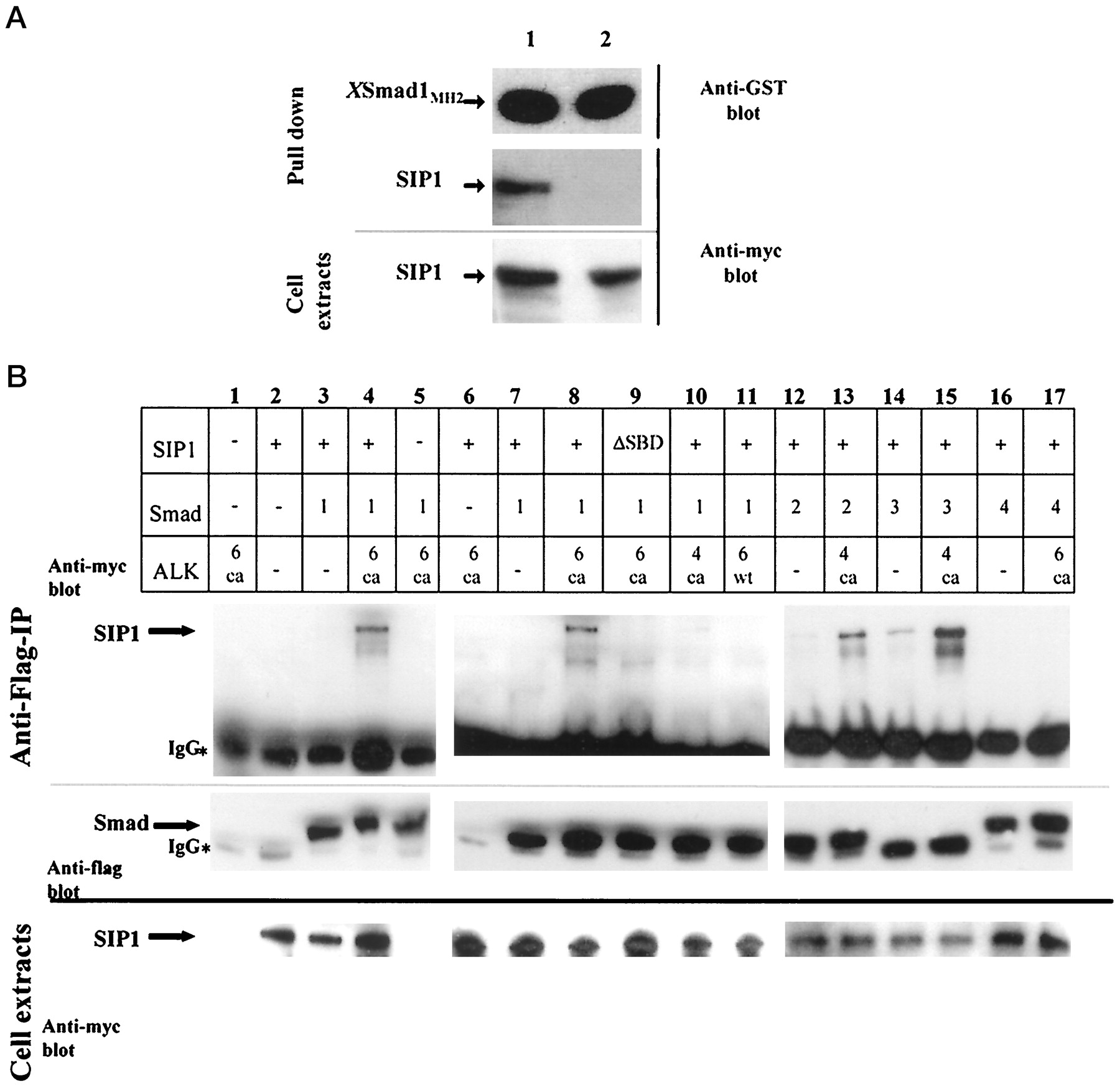
FIG. 5. A, interaction of SIP1 with the MH2 domain of XSmad1 in mammalian cells. An expression construct encoding a fusion between GST and
the MH2 domain of XSmad1 was transfected in COS1 cells together with expression constructs for Myc-tagged SIP1. As shown by immunoblotting
of pulled-down material from cell extracts, SIP1 specifically interacted with the GST-XSmad1 fusion protein (middle panel, lane 1), whereas
deletion of the SBD51 disrupted the interaction (middle panel, lane 2). Comparable affinity purification of the GST-fusion protein and equal
expression of SIP1 were confirmed by immunoblotting of the pulled down material using polyclonal anti-GST antibody (upper panel) and of total
cell extracts using monoclonal anti-myc antibody (lower panel), respectively. B, ligand-dependent interaction of SIP1 with full-length Smads in
mammalian cells. Lanes 1–5, lanes 6–11, and lanes 11–17 contain data from three independent experiments. HEK293T cells were transiently
transfected with various combinations of expression constructs encoding Myc-tagged SIP1, Flag-tagged Smads, and type I receptors, as indicated.
Cell lysates were immunoprecipitated with anti-Flag antibodies, and the precipitated proteins were visualized by SDS-polyacrylamide gel
electrophoresis and immunoblotting using anti-Myc (upper panel) or anti-Flag (middle panel) antibodies. The middle panel shows the comparable
immunoprecipitations of Flag-tagged Smads in each experiment, whereas the lower panel shows immunoblotting of total cell extracts using
anti-Myc antibody, to confirm comparable expression of SIP1. *, indicates the heavy chain of the anti-Flag antibody used in the immunoprecipitations;
ca, constitutively active; wt, wild type; DSBD indicates SIP1 in which the 51-amino acids-long SBD was deleted. Image published in: Verschueren K et al. (1999) Copyright © 1999. Image reproduced with permission of the Publisher. Larger Image Printer Friendly View |
