XB-IMG-140031
Xenbase Image ID: 140031
|
|
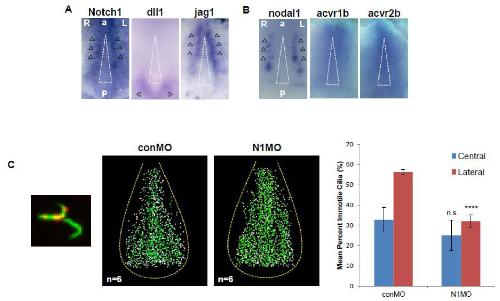
Figure S1. Notch signaling regulates the fate of cilia at the border of the GRP, Related to Figure 1.
A. Expression of Notch pathway components Notch1, dll1 and jag1 at the GRP at stage 13/14, analyzed by WISH. B. Expression of TGF- pathway components nodal1, acvr1b and acvr2b at the GRP at stage 13/14, analyzed by WISH. Black arrowheads indicate normal expression of Notch1, dll1, jag1 and nodal1 expression. acvr1b and acvr2b expression were detected in the GRP. A white triangle indicates the central region of the GRP. C. The ratio of motile to immotile cilia increased at the border of Notch1-depleted GRPs. The left panel shows representative LRD-positive and LRD-negative cilia. Acetylated -tubulin or LRD is represented by green or red signal, respectively. The two panels in the middle, conMO injected and N1MO injected, show the distribution of cilia subtypes at the GRP. These panels represent data from 6 samples, overlaid in Photoshop. Green or white dots represent motile or immotile cilia, respectively. The right graph shows the quantification of immotile cilia in the lateral area and the central area of control or Notch1-depleted GRPs. Lateral or central regions were delineated using previously described methods (Boskovski et al., 2013). **** ≤ 0.00005, n.s.: not significant. Image published in: Tözser J et al. (2015) Copyright © 2015. Image reproduced with permission of the Publisher and the copyright holder. This is an Open Access article distributed under the terms of the Creative Commons Attribution License. Larger Image Printer Friendly View |
