XB-IMG-148940
Xenbase Image ID: 148940
|
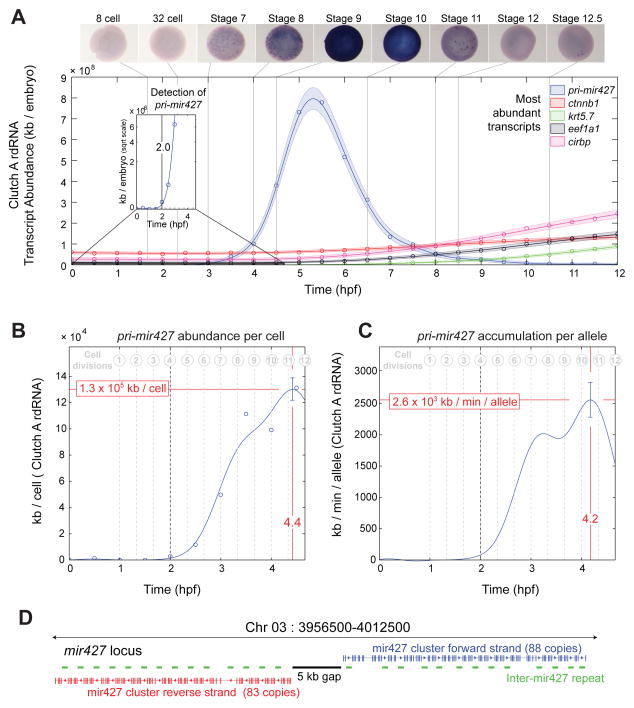
Accumulation kinetics of pri-mir427 per allele in Clutch A rdRNA, see also Fig S7
A â pri-mir427 expression over first 12 hpf in rdRNA data and in situ hybridizations. pri-mir427 show ubiquitous expression matching RNA-seq temporal data. Examples of most abundant transcripts shown for comparison. Inset: first detection of pri-mir427 above detection limit at 2.0 hpf (8â16 cell transition).
B â pri-mir427 abundance in kb per cell. Line marks Gaussian process median, error bar is 95% CI (Supplemental Experimental Procedures). Peak per cell abundance occurs after 11th division at 4.4 hpf.
C â pri-mir427 accumulation rate in kb/min/allele with time. Line is median of differential of Gaussian Process, error bar gives 95% CI (Supplemental Experimental Procedures). Peak accumulation rate is achieved at 4.2 hpf just after 10th cell division.
D â Organization of pri-mir427 locus on X. tropicalis Chromosome 3 (X. tropicalis v8 genome, Supplemental Experimental Procedures). Clusters of mir427 hairpins appear in tandem with a repeated sequence arranged symmetrically on opposite strands around a gap in the genome assembly. Image published in: Owens ND et al. (2016) Copyright © 2016. Image reproduced with permission of the Publisher and the copyright holder. This is an Open Access article distributed under the terms of the Creative Commons Attribution License. Larger Image Printer Friendly View |
