XB-IMG-150820
Xenbase Image ID: 150820
|
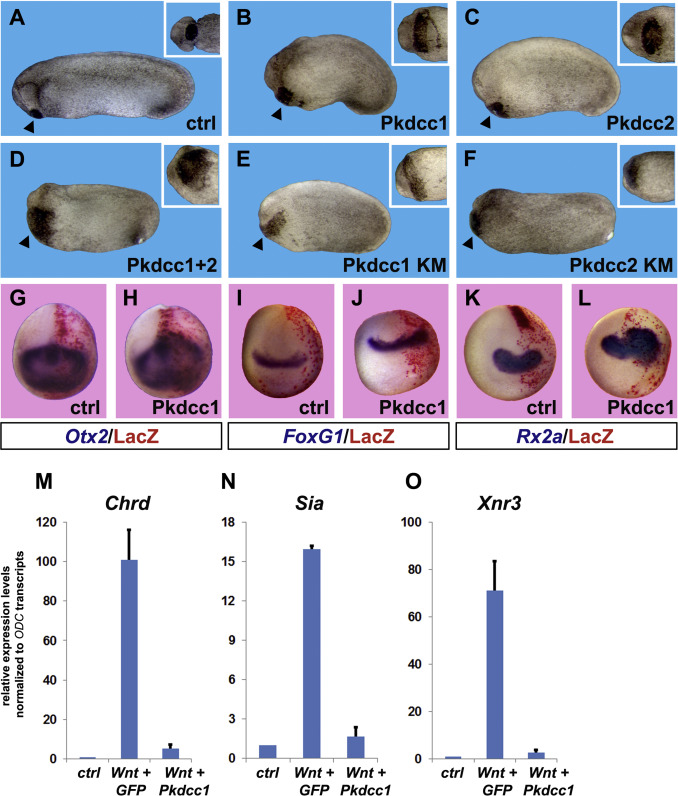
Fig. 6. Overexpression of Pkdcc enlarges anterior tissues and inhibits canonical Wnt signaling. (AâC) Overexpression of 1 ng of Pkdcc1 or Pkdcc2 mRNA caused expansion of anterior structures such as the cement gland (see insets). Percent of embryos showing enlarged cement gland and head were 0% n=60 for controls; 81% n=54 for Pkdcc1; 84% n=38 for Pkdcc2. (D) This effect is synergistic, since injection of 500 pg each of Pkddc1 and 2 caused an even greater enlargement of the cement gland (98% of the embryos showed this phenotype, n=50). (EâF) Kinase mutant (KM) forms of Pkdcc1 and 2 also caused an increase of cement gland tissues, suggesting that the kinase activity is dispensable for Pkdcc-mediated anteriorizing effects. Percent of embryos showing enlarged cement glands were: 82% n=27 for Pkdcc1-KM; 83% n=24 for Pkdcc2-KM. All embryos are shown from lateral views, insets show ventral view of the cement gland from the same embryos. Arrowheads point to cement gland. (GâL) Whole-mount in situ hybridizations for anterior neural markers show that Pkdcc1 mRNA overexpression expands Otx2,FoxG1 (also known as BF1) and Rx2a. The injected side was identified through Red Gal staining of the lineage tracer nuclear Lacz (nLacz) coinjected with Pkdcc1. Control embryos were injected with nLacz only. Percentage of Pkdcc1-injected embryos showing expanded neural markers: Otx2 66% n=27; FoxG1 76% n=47; Rx2a 92% n=38. (MâO) Pkdcc1 abolishes canonical Wnt signaling. Quantitative RT-PCR analysis performed on animal cap explants co-injected with xWnt8 and Pkdcc1 mRNAs shows that Pkdcc1 greatly reduces induction of canonical Wnt target genes Chordin (Chrd), Siamois (Sia) and Nodal related 3 (Xnr3). GFP mRNA was used as a negative control. Transcript levels of the housekeeping gene ODC were used for normalization. Error bars indicate standard deviations derived from three independent experiments. Image published in: Ding Y et al. (2017) Copyright © 2017. Image reproduced with permission of the Publisher, Elsevier B. V.
Image source: Published Larger Image Printer Friendly View |
