XB-IMG-75180
Xenbase Image ID: 75180
|
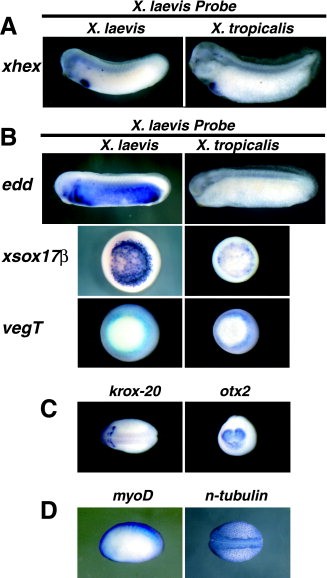
Figure 4. A,B: In situ hybridizations were performed for endodermally expressed genes using Xenopus laevis probes in X. laevis embryos (left column) and X. tropicalis embryos (right column). X. laevis xhex probe works well on both species (A), whereas most of the X. laevis endodermal probes tested worked poorly on X. tropicalis embryos (B). For xhex and endodermin (edd), lateral views (anterior to the left) of stage 28â29 embryos are shown. For xsox17β and vegT, vegetal views of midgastrula stage embryos are shown. For all of the X. laevis probes used in this study, a run was tried without RNase to determine which X. laevis probes could be enhanced in X. tropicalis embryos. Most probes showed no enhancement (not shown). Some showed better staining (C), whereas others showed nonspecific staining (D) when the RNase treatment was removed from the protocol. Dorsal views (anterior to the left) of stage 20 embryos are shown for krox-20 and n-tubulin. A lateral view (anterior to the left) of stage 20 embryos are shown for myoD. An anterior view (dorsal up) of a stage 20 embryo is shown for otx2.Download figure to PowerPoint Image published in: Khokha MK et al. (2002) Copyright © 2002. Image reproduced with permission of the Publisher, John Wiley & Sons. Larger Image Printer Friendly View |
