XB-IMG-84073
Xenbase Image ID: 84073
|
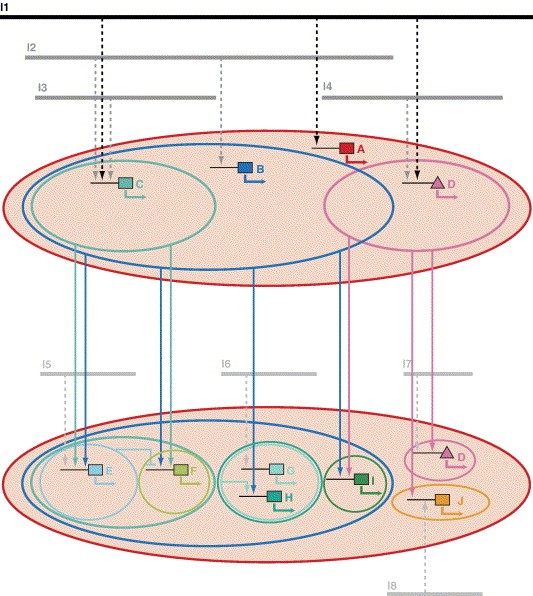
Fig. 7. Combinatorial multistep model for placode induction and specification. Expression of various transcription factor genes involved in placode specification (colored rectangles and triangles, A–J) may be regulated by particular combinations of upstream factors including inducing signaling molecules (I1–I8) as well as other transcription factors, which bind to their cis-regulatory region. Upstream factors may be activating (arrows; broken arrows are used for inducing signals because these act indirectly via transcription factors downstream of signaling cascades) or repressing (bars). In the hypothetical case illustrated, upstream factors are assumed to be singly necessary but only jointly sufficient for activation of the downstream gene. The expression domain of a transcription factor (enclosed by colored lines; the red area represents the panplacodal primordium) is determined by the spatial extent of the expression domains of inducers and transcription factors acting upstream (indicated by black and gray lines for inducers and colored ellipses for transcription factors). The expression of transcription factors specific for individual placodes (lower panel) depends on panplacodal (black), multiplacodal (dark gray), and placode-specific (light gray) inducers, which may act either directly or indirectly (i.e. mediated by other transcription factors with a panplacodal, multiplacodal, or placode-specific distribution). Some transcription factors (as illustrated for D), which show placode-specific expression at later stages, may initially be more broadly expressed but then become restricted because they require input from more localized inducers for sustained activation. This model implies that different placodes share some but not all of the inducers and transcription factors involved in their specification. Moreover, it proposes that at early stages of placode specification (upper panel) there are nested and partially overlapping regions differentially biased for development of different sets of placodes. Multiplacodal bias is introduced by transcription factors (such as B–D) which cover multiple prospective placodes and activate various placode-specific transcription factors, without being by themselves sufficient for their activation. Panplacodally expressed transcription factors such as A may, in addition, activate genes involved in the regulation of generic placodal processes such as proliferation, cell shape changes, and neurogenesis (not shown). Image published in: Schlosser G (2006) Copyright © 2006. Image reproduced with permission of the Publisher, Elsevier B. V. Larger Image Printer Friendly View |
