XB-IMG-84486
Xenbase Image ID: 84486
|
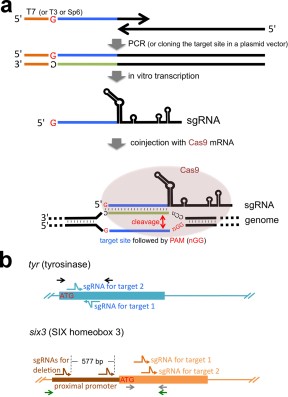
Figure 1. Strategy for CRISPR/Cas-mediated genome modification. (a) Schematic representation of experimental procedure. We found that the conventional plasmid subcloning method for making sgRNA templates was time-consuming and inconvenient; instead all sgRNAs used in this study are transcribed in vitro from double-stranded DNA templates that were made by PCR (except the sgRNA for tyr target site 1, which used a cloned template, pDR274-Xt-tyr). This PCR strategy uses a 5′ oligonucleotide (primer) that begins with the T7 promoter (shown as an orange line in this schematic; alternatively, T3 or SP6 promoters could be used) and contains the genomic target sequence (shown as a blue line in this schematic; note the genomic target sequence must begin with a G for proper transcriptional initiation using the T7 promoter) and a 3′ oligonucleotide (primer) that partly overlaps the 5′ primer and contains the sgRNA backbone sequence required for proper folding of sgRNAs. This rapid, easy way to make sgRNA templates by PCR takes less than two days to produce sgRNAs to inject once oligonucleotides are received. During the preparation of this manuscript, we have noticed that similar strategies have also been reported [e.g., (Bassett et al., 2013)] and thus this method has general versatility. The resulting sgRNAs were coinjected with Cas9 mRNA into one cell-stage Xenopus embryos. The bottom part of the scheme shows how sgRNA and Cas9 work to cleave the target site in the genome. Briefly, the sgRNA forms a complex with the Cas9 protein and identifies the target site via complementary base-pairing. The Cas9 nuclease subsequently cleaves the genomic DNA at the target site, just upstream of the PAM sequence. (b) Targeting strategy for the two genes described in this study. Both tyr and six3 genes were targeted in exon 1 to cause frame-shifts after the translation initiation codon (ATG). Two independent sgRNAs were tested for both genes. The six3 gene was furthermore targeted in the proximal promoter region with two sgRNAs simultaneously to cause deletion of the promoter region. Bent arrows (blue for tyr, orange for six3 exons and brown for the six3 promoter) show sgRNA targets. Arrows (black for tyr, green and grey for six3) indicate genomic PCR primers for mutation analyses. Note that the 3' primer for the six3 gene is common for both sets of PCR reactions but colored differently for clarity. Drawings are not to scale. Image published in: Nakayama T et al. (2013) Copyright © 2013. Image reproduced with permission of the Publisher, John Wiley & Sons. Larger Image Printer Friendly View |
