A celebration of the life and genius of Yoshiki Sasai (1962-2014): An obituary
by Enrique Amaya
August 5th 2014 will forever mark one of the saddest days in the fields of Xenopus embryology, developmental and stem cell biology, and regenerative medicine, as on this fateful day, the life of one of the greatest minds in the biomedical sciences came to a tragic and premature end. Here, I commemorate the extraordinary life and achievements of our friend, mentor and collaborator, Yoshiki Sasai.
The formative years
Yoshiki Sasai was born on 5th of March 1962 in Hyogo, Japan. After obtaining a medical degree from Kyoto University Medical School in 1986, Yoshiki spent the next two years doing an internship in general and emergency medicine. However, after his medical internships, he returned to Kyoto University Medical School and entered their PhD programme, as a career in academic research beckoned versus one spent in the clinics. He joined the lab of Shigetada Nakanishi and at this time his lifelong interest in brain development was born. As part of his PhD thesis he cloned and partially characterised the first mammalian orthologues of the Drosophila hairy and enhancer of split gene; namely Hes1, Hes2, Hes3 and Hes4 (Sasai et al., 1992). Hes genes have subsequently been shown to display oscillatory expression dynamics and play essential roles during both neurogenesis and somitogenesis (reviewed in (Kageyama et al., 2007).
After completing his PhD in 1993, Yoshiki moved to California to join the laboratory of Eddy De Robertis at UCLA. Here he was introduced to classical and modern molecular embryology using Xenopus as an experimental system. Although he only spent three years in Eddy’s lab, his post-doctoral time at UCLA was formative, incredibly productive and highly influential in his intellectual development. The initial aim of Yoshiki’s research was to identify the signalling molecules that mediate the activities of Spemann’s organizer. Eddy’s lab had previously identified a gene expressed in the organizer, goosecoid (gsc), which encoded a homeobox containing transcription factor and exhibited organiser activity when misexpressed in Xenopus embryos (Cho et al., 1991). A key challenge that remained was to identify the critical signalling (i.e. secreted) factors that acted downstream of gsc, which directly (as opposed to indirectly) mediated the activity of the organiser. This was the question that Yoshiki set out to answer.
Although the nature of the signalling molecules produced by the Spemann’s organiser had eluded scientists for over fifty years, within a year of joining Eddy’s lab Yoshiki identified Chordin, as one of the critical signalling molecules produced by the Spemann’s organiser (Sasai et al., 1994). The gene, chordin (chd), encoded a secreted molecule, it was expressed in the right place and at the right time (dorsal lip of the blastopore and subsequently the notochord; see Figure), and it exhibited the right activity (i.e. it could induce a secondary axis, when misexpressed in the ventral side of the Xenopus embryo), and could thus recapitulate the actions of Spemann’s organiser. A year later, in 1995, Yoshiki went on to show that Chordin could directly induce neuroectoderm from naïve, competent embryonic ectoderm by antagonising BMP4 signalling (Sasai et al., 1995). In this way, the identification of Chordin, by Yoshiki, alongside the identification of other signalling proteins with similar activity (i.e., Noggin and Follistatin) ended one of the remaining Holy Grails of embryology; namely, the identity of the secreted molecules responsible for neural induction.
However, Yoshiki’s prolific tenure in Eddy’s lab did not end there. Eddy and Yoshiki devoured much of the classical scientific literature examining embryology and they delved into comparative studies, a field now termed “evolutionary developmental biology” or evo-devo. They noted that the ortholog of chd in Drosophila was a gene called, short gastrulation (sog). This gene had previously been shown to antagonise the function of dpp, the Drosophila orthologue of BMP4, where it was similarly involved in the establishment of the dorsoventral (D/V) axis in the fruit fly embryo (Holley et al., 1995). However, there was one fascinating difference between the functions of these gene toolkits in Drosophila versus Xenopus. While Chordin-mediated antagonism of BMP4 helped establish the dorsal domain in the vertebrate embryo, Sog mediated antagonisms of Dpp helped establish the ventral domain of the fruit fly embryo. Thus the chd/sog/bmp4/dpp gene toolkit was ancestral and likely to have played a key role in D/V patterning in the embryo of the common ancestor of arthropods and chordates, yet intriguingly, there had been an inversion of D/V axis at some point after this common ancestor in one of the lineages, an idea that had been previously put forward by E. Geoffrey Saint-Hilaire in 1822 (De Robertis and Sasai, 1996). Furthermore, based partly on this evidence, Eddy and Yoshiki proposed a name for the common ancestor of all deuterstomes and protostomes, namely the Urbilateria (i.e. primitive bilateral common ancestor), which existed around 600 million years ago, and they went on to propose five developmental gene programmes exhibited by this primitive common ancestor (De Robertis and Sasai, 1996).
The consolidation years
After spending three years in California, Yoshiki returned to Japan in 1996 to take up an Associate Professor position in his alma mater, Kyoto University, where he quickly progressed to full Professor within two years. Then in 2000 he became one of the founding faculty members at the RIKEN Center for Developmental Biology, in Kobe, where he was Director of the Laboratory of Neurogenesis and Organogenesis. After returning to Japan, he continued to exploit Xenopus for its strengths in molecular embryological approaches, focusing primarily on the mechanisms of neural induction and neurogenesis. However, during this time, he lamented that studies on neural induction in mammals was significantly behind those using Xenopus and he felt that this was due to the lack of a good experimental system in mammals for in vitro studies of neural induction. Thus, his vision was to transform mammalian neural induction and neurogenesis studies such that one could explore these in ways similar to those he had become so accustomed to using Xenopus. In particular, his view was that one might be able to use pluripotent mouse embryonic stem (mES) cells in very much the same way that Xenopus developmental biologists had been using pluripotent animal cap cells to investigate the mechanisms of neural induction for many years prior. This vision was clearly evident in one of his earliest articles using mES cells in which he and colleagues generated midbrain dopaminergic neurons in vitro (Kawasaki et al., 2000). In the introduction of this paper he wrote:
“During the last decade, much progress has been made in the molecular understanding of early neural differentiation in Xenopus. … By contrast, relatively little is known about regulatory factors in mammalian neural induction. One main reason for this is that good experimental systems for in vitro neural differentiation are still lacking in mice (i.e., something comparable to the animal cap assay commonly used in Xenopus studies).”
He then went on to propose that embryonic stem cells could provide that transition and presented data showing the efficient in vitro derivation of dopaminergic neurons using serum free conditions from mES cells, and then he discussed the possible future therapeutic potential of these cells in the treatment of Parkinson’s disease.
After this paper, he made quick and remarkable progress in exploiting mES cells for the dual purpose of understanding the mechanisms that drive the development of the mammalian nervous system, but also as a means of generating neural cell types with potential translational and therapeutic applications in the treatment of human diseases. For example, he identified conditions to derive telencephalic precursors from mES cells (Watanabe et al., 2005), and furthermore, he optimized ways to generate various specific neuronal subtypes, such as Purkinje cells, from mES cells (Muguruma et al., 2010). However, the most remarkable advance that Yoshiki made was the identification of conditions that would permit the in vitro self-assembly and differentiation of various neural structures using 3-D cultures of embryonic stem cells. Firstly he published conditions that would result in polarized cortical tissues. (Eiraku et al., 2008). More recently, however, Yoshiki and colleagues modified their 3-D culture conditions in order to allow the self-organization and morphogenesis of fully formed and differentiated eyes and anterior pituitaries from mES cells (Eiraku et al., 2011; Suga et al., 2011) (see also Figure). The potential translational and therapeutic relevance of these findings in the bourgeoning field of regenerative medicine are undeniable. Although much of these ground-breaking studies were done using mES cells, Yoshiki also made significant contributions to the field of human embryonic stem (hES) cells. Notable amongst these was the identification of conditions that would permit the long term survival of hES cells in vitro, which facilitated the exploitation of these cells for both basic studies of human development, and importantly, their therapeutic use in the future (Watanabe et al., 2007). Yoshiki was then able to modify his 3-D in vitro culture conditions with hES cells in order to facilitate the self-organization and generation of human eyes in vitro (Nakano et al., 2012).
The repercussions of these findings are tremendous; for example, the ES-cell technologies developed by Yoshiki can now be used to generate human optic tissues from patient derived human iPS cells in order to gain better insight into the aetiology of various inherited retinal dystrophies and other congenital eye diseases. Furthermore, his methods can now be exploited for the derivation of patient-specific tissues and organs (rather than cells), useful for transplantation into patients. The future of regenerative medicine and its applications to treat human diseases is bright, and Yoshiki will forever be one of the giants upon whose shoulders those who follow will stand, as patient-specific tissue and organ replacement therapies become a reality. This will be the legacy of Yoshiki’s brilliance and vision. He was not only one who could foresee the future of regenerative medicine clearly, but he also was one who drove the field forward like no other. His insight and vision was clearly evident in a commentary he published in 2008 in the journal, Growth, Development and Differentiation, entitled “Bridging the gap from frog research to human therapy: A tale of neural differentiation in Xenopus animal caps and human pluripotent cells” (Sasai et al., 2008). Intriguingly, although he wrote this commentary with three other co-authors, the article was written in the first tense, and it expounds his own vision succinctly and clearly. In the conclusion of this commentary Yoshiki asks: “So is basic embryological research no longer necessary?” He answers his own question by stating that:
“There is still plenty of room for a substantial contribution from basic embryological research in many senses. On top of this, in terms of regenerative medicine, cell therapy with human ES/iPS cells is just a beginning. A cell is a cell and not an organ. In the near future, basic embryological studies on organogenesis, I believe, will provide key information and technologies that will elevate the level of human ES- and iPS-cell-derived therapeutics to a new dimension”
And that will forever be his legacy.
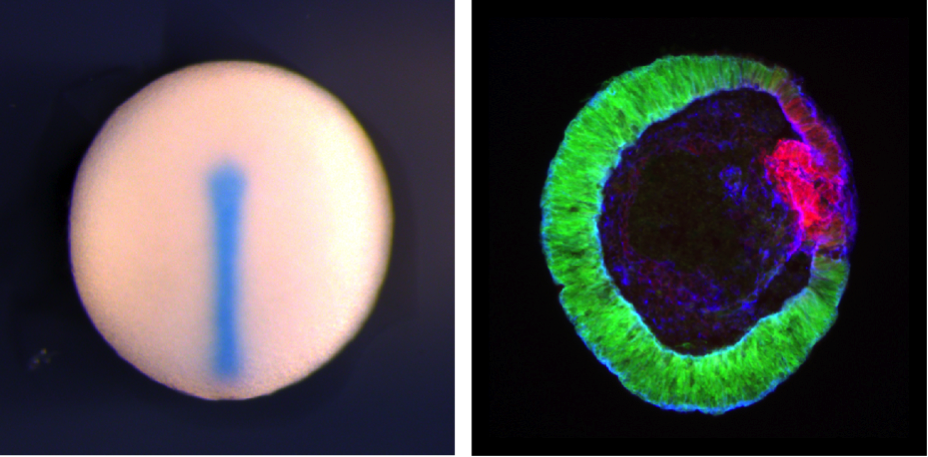
Left, midline of St. 20 Xenopus embryo showing chordin expression using in situ hybridization (by Siwei Zhang). Right, Cryosection of mouse Rx:GFP ES cell-derived optic tissue generated using "SFEBq" (by Nick Love). Rx:GFP signal shows neural retinal tissue, Red show Mitf expression, a marker for retinal pigmental epithelium. Blue shows actin.
References
Cho, K.W., B. Blumberg, H. Steinbeisser, and E.M. De Robertis. 1991. Molecular nature of Spemann's organizer: the role of the Xenopus homeobox gene goosecoid. Cell. 67:1111-1120.
De Robertis, E.M., and Y. Sasai. 1996. A common plan for dorsoventral patterning in Bilateria. Nature. 380:37-40.
Eiraku, M., N. Takata, H. Ishibashi, M. Kawada, E. Sakakura, S. Okuda, K. Sekiguchi, T. Adachi, and Y. Sasai. 2011. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature. 472:51-56.
Eiraku, M., K. Watanabe, M. Matsuo-Takasaki, M. Kawada, S. Yonemura, M. Matsumura, T. Wataya, A. Nishiyama, K. Muguruma, and Y. Sasai. 2008. Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals. Cell stem cell. 3:519-532.
Holley, S.A., P.D. Jackson, Y. Sasai, B. Lu, E.M. De Robertis, F.M. Hoffmann, and E.L. Ferguson. 1995. A conserved system for dorsal-ventral patterning in insects and vertebrates involving sog and chordin. Nature. 376:249-253.
Kageyama, R., T. Ohtsuka, and T. Kobayashi. 2007. The Hes gene family: repressors and oscillators that orchestrate embryogenesis. Development. 134:1243-1251.
Kawasaki, H., K. Mizuseki, S. Nishikawa, S. Kaneko, Y. Kuwana, S. Nakanishi, S.I. Nishikawa, and Y. Sasai. 2000. Induction of midbrain dopaminergic neurons from ES cells by stromal cell-derived inducing activity. Neuron. 28:31-40.
Muguruma, K., A. Nishiyama, Y. Ono, H. Miyawaki, E. Mizuhara, S. Hori, A. Kakizuka, K. Obata, Y. Yanagawa, T. Hirano, and Y. Sasai. 2010. Ontogeny-recapitulating generation and tissue integration of ES cell-derived Purkinje cells. Nature neuroscience. 13:1171-1180.
Nakano, T., S. Ando, N. Takata, M. Kawada, K. Muguruma, K. Sekiguchi, K. Saito, S. Yonemura, M. Eiraku, and Y. Sasai. 2012. Self-formation of optic cups and storable stratified neural retina from human ESCs. Cell stem cell. 10:771-785.
Sasai, Y., R. Kageyama, Y. Tagawa, R. Shigemoto, and S. Nakanishi. 1992. Two mammalian helix-loop-helix factors structurally related to Drosophila hairy and Enhancer of split. Genes & development. 6:2620-2634.
Sasai, Y., B. Lu, H. Steinbeisser, and E.M. De Robertis. 1995. Regulation of neural induction by the Chd and Bmp-4 antagonistic patterning signals in Xenopus. Nature. 376:333-336.
Sasai, Y., B. Lu, H. Steinbeisser, D. Geissert, L.K. Gont, and E.M. De Robertis. 1994. Xenopus chordin: a novel dorsalizing factor activated by organizer-specific homeobox genes. Cell. 79:779-790.
Sasai, Y., M. Ogushi, T. Nagase, and S. Ando. 2008. Bridging the gap from frog research to human therapy: a tale of neural differentiation in Xenopus animal caps and human pluripotent cells. Development, growth & differentiation. 50 Suppl 1:S47-55.
Suga, H., T. Kadoshima, M. Minaguchi, M. Ohgushi, M. Soen, T. Nakano, N. Takata, T. Wataya, K. Muguruma, H. Miyoshi, S. Yonemura, Y. Oiso, and Y. Sasai. 2011. Self-formation of functional adenohypophysis in three-dimensional culture. Nature. 480:57-62.
Watanabe, K., D. Kamiya, A. Nishiyama, T. Katayama, S. Nozaki, H. Kawasaki, Y. Watanabe, K. Mizuseki, and Y. Sasai. 2005. Directed differentiation of telencephalic precursors from embryonic stem cells. Nature neuroscience. 8:288-296.
Watanabe, K., M. Ueno, D. Kamiya, A. Nishiyama, M. Matsumura, T. Wataya, J.B. Takahashi, S. Nishikawa, S. Nishikawa, K. Muguruma, and Y. Sasai. 2007. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nature biotechnology. 25:681-686.
Last Updated: 2014-08-25
