Comprehensive analyses of hox gene expression in Xenopus laevis embryos and adult tissues
Kondo M, Yamamoto T, Takahashi S, Taira M.
Dev Growth Differ. August 1, 2017; 59 (6): 526-539.
Click here to view the article at Development, Growth, and Differentiation.
Click here to view the article on Pubmed.
Click here to view the article on Xenbase.
Abstract
From whole genome sequencing of an allotetraploid frog, Xenopus laevis, two homeologous sets (L and S) of four Hox clusters A through D (HoxA.L/S, HoxB.L/S, HoxC.L/S, and HoxD.L/S) and 13 paralogous groups (PGs) with 76 genes were identified, allowing us to carry out the first comprehensive analyses of hox gene expression in vertebrates. Expression of all hox genes during development and in adult tissues was analyzed by RNA-sequencing. The expression levels of most hox genes were similar between homeologs, but in some pairs, large differences were observed and several of these were confirmed by RT-PCR and whole mount in situ hybridization experiments. These results indicate that subfunctionalization of hox genes may have occurred since allotetraploidization. Furthermore, comprehensive analysis of hox gene expression during early development did not agree with the hypothesis of temporal collinearity especially in genes belonging to PG2 to PG10.

Fig. 1. The Hox clusters of Xenopus laevis and X. tropicalis. (A) Schematic repre-sentations of the four and eight Hoxclusters of X. tropicalis and X.laevis, respectively. hox genes of 13 paralogousgroups (PGs) are present. even-skippedhomeobox (evx) genes are present onHoxA and HoxD clusters. The four pairsof homeologous Hox clusters of X. laevisconsist of clusters from its two subgenomes, and are designated as L and S.hoxb2p.L is a pseudogene (Session et al. 2016). Positions of micro RNA genesmir10a (blue arrowheads) and mir10b (redarrowheads) are also shown. (B) Positions of Hox clusters on X. laevis chromo-somes. Hox clusters are present on corre-sponding chromosomes of the twosubgenomes at corresponding positions.

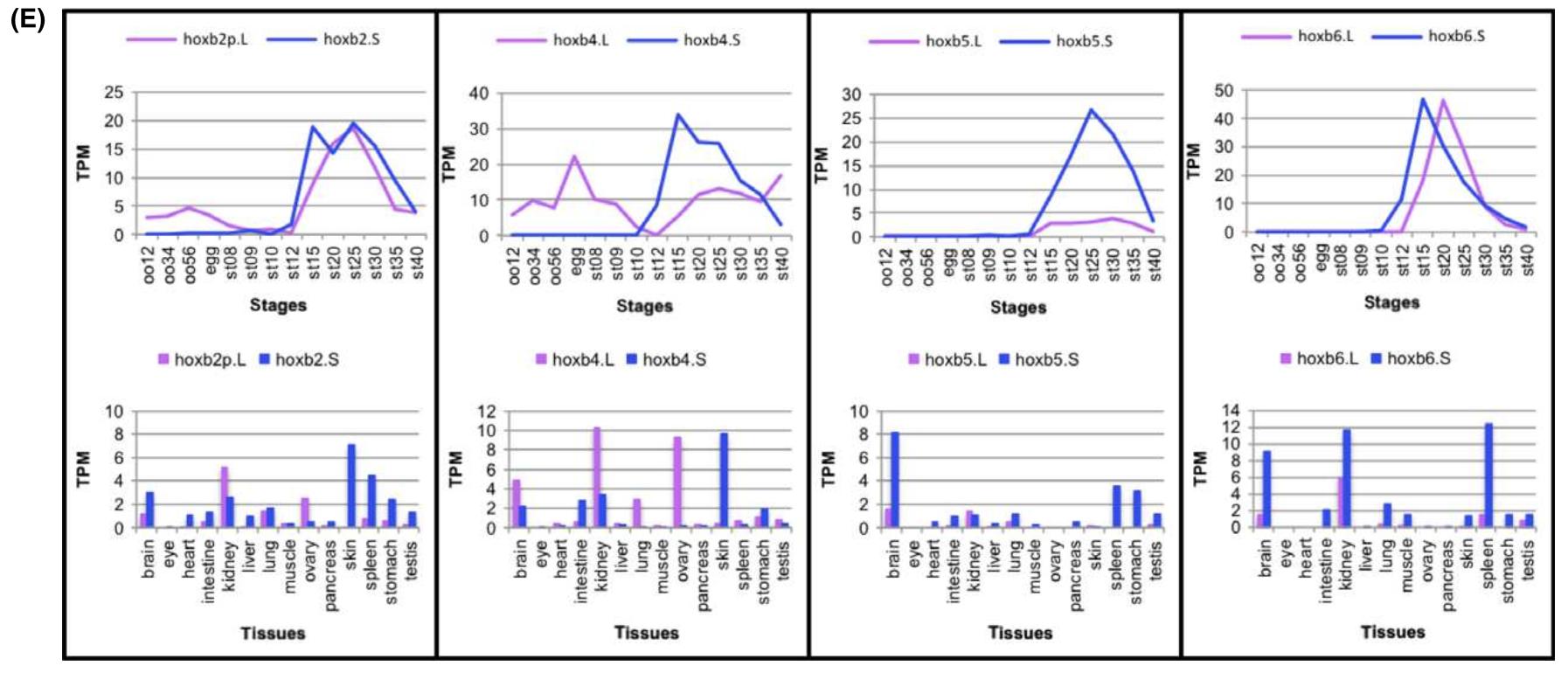
Fig. 2. Expression of hox genes. (A–D) The expression of all 76 hox genes from four pairs of Hox clusters in X. laevis was analysed byRNA-seq analyses during developmental stages (stage (st) 8–40). The average values of TPM from two clutches are plotted for eachgene. Only the PG number is indicated where there is no corresponding gene in that cluster. Between some homeologs, a large difference in expression levels are observed (for example, hoxb5.L and hoxb5.S [arrows]). Note that hoxb4.L is expressed maternally (asterisk). (E) Expression profiles during development and in adult tissues of hoxb2, hoxb4, hoxb5, and hoxb6 homeologs. Average TPMvalues are plotted. oo12: oocyte stages I and II; oo34: oocyte stages III and IV; oo56: oocyte stages V and VI; st: stage.



Fig. 4. Temporal expression of hox genes during developmental stages 8 to 40. (A) Expression levels of 50 genes that had TPM more than 5.0 at some point was normalized so that the peak TPM is 1. By clustering analysis, genes were grouped into eight “expressiongroups” (ExpG). hoxb4.L was exceptional, since the egg showed the highest value, but to observe zygotic expression, it was normalized to the value at stage 40. Nevertheless, because of its unique pattern, this was not included in any ExpG. (B) Expression profies of eachExpG. The patterns are mostly overlapping. (C) Summary of expression profiles. Four of the six PG1 (labial) genes are categorized inExpG1 (the other two are ExpG4), and expressed early, and PG11 to 13 genes in the posterior group are expressed late (ExpG8) or notuntil after stage 40, whereas the pb, central, and posterior PGs 9 and 10 showed no temporal collinearity. Asterisks indicate homeologsthat belong to the same ExpG.
Adapted with permission from Wiley: Kondo et al. (2017). Comprehensive analyses of hox gene expression in Xenopus laevis embryos and adult tissues. Dev Growth Differ. 2017 Aug;59(6):526-539. DOI: 10.1111/dgd.12382. Copyright 2017.
