XB-IMG-148946
Xenbase Image ID: 148946
|
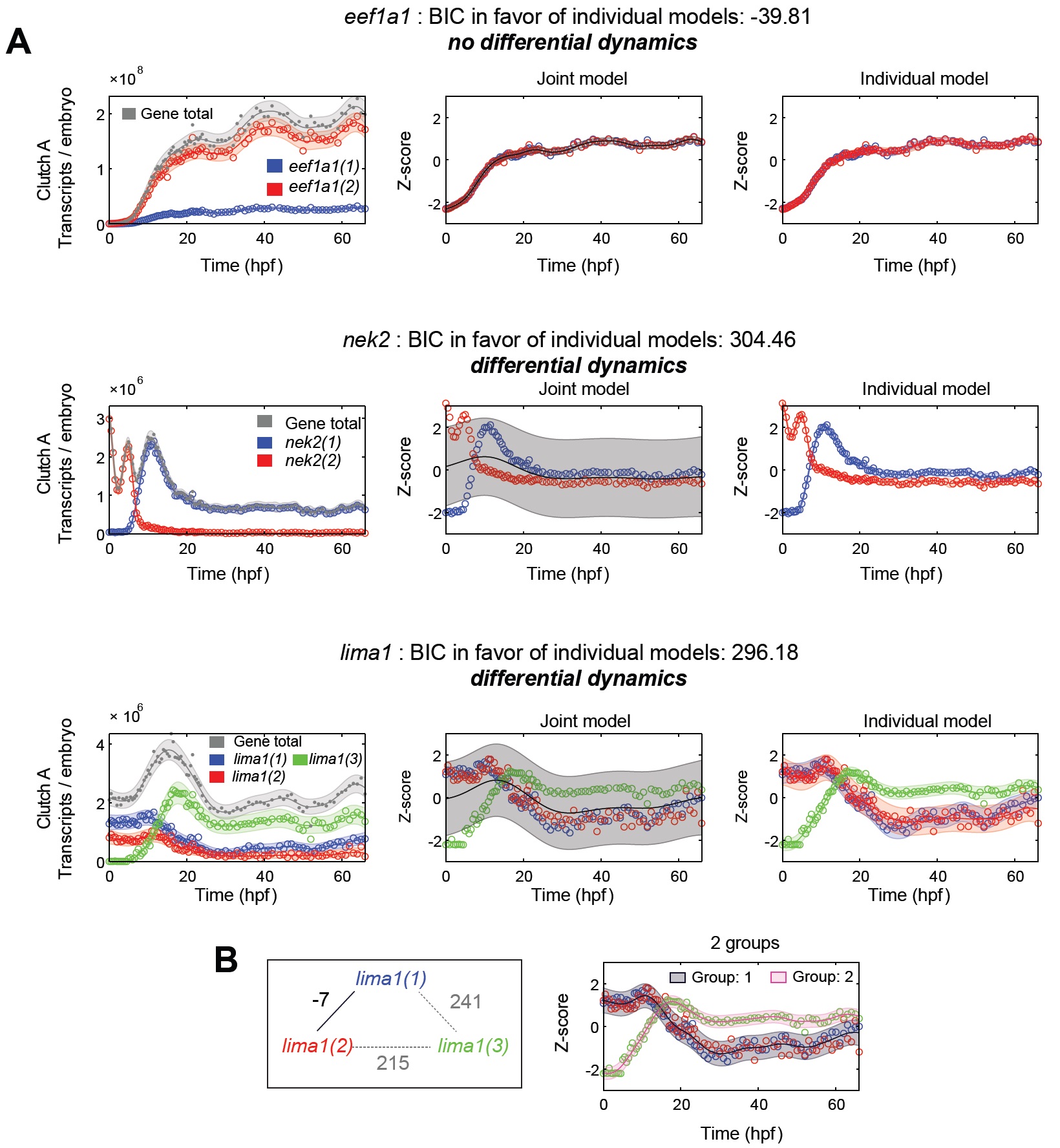
Fig S5 – Differential Isoform Dynamics in Clutch A polyA+, related to Fig 3.
(A) Explanation and detection of differential isoform dynamics. Left: isoform expression in transcripts/embryo. Center: Joint
model of all isoforms assumed to have the same expression on z-normalized scale (see Supplemental Experimental Methods).
Right: Models of all isoforms considered separately. If Bayesian Information Criterion (BIC) favors the joint model, the gene has
differential abundance, whereas if the BIC favors the individual models a gene has differential dynamics. eef1a1 has differential
abundance, nek2 and lima1 have differential dynamics.
(B) Differential dynamics of groups of isoforms, BIC graph (left) between all lima1 isoforms. lima2(1) and lima2(2) have differential
abundance, and they have differential dynamics with lima2(3) defining two groups of isoforms (right). Image published in: Owens ND et al. (2016) Copyright © 2016. Image reproduced with permission of the Publisher and the copyright holder. This is an Open Access article distributed under the terms of the Creative Commons Attribution License. Larger Image Printer Friendly View |
