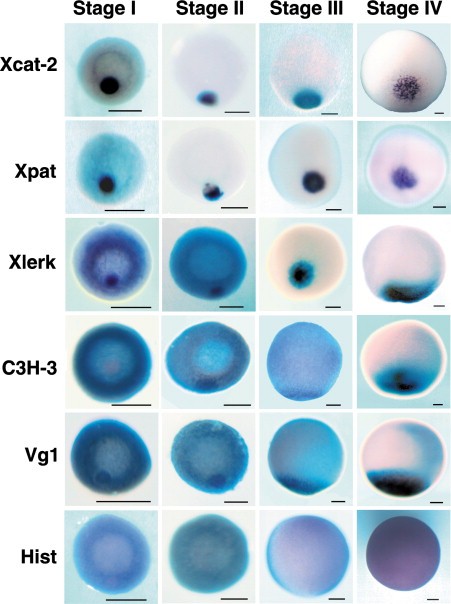
Figure 2. In Situ Hybridization of Newly Discovered Localized mRNAsIn situ hybridization toXcat-2 mRNA and Xpat mRNAs demonstrates examples of RNAs that localize via the early pathway. Note that most of the RNA is associated with the mitochondrial cloud in stage I oocytes, with little background in the rest of the cytoplasm. By stage II, these RNAs have migrated with the cloud to the vegetal cortex, and there is no detectable RNA in the remaining cytoplasm. Xlerk, which was identified by computational screening of Xenopus 3â² UTRs for highly significant clusters of CAC-containing motifs, follows the intermediate pathway in which localization to the cloud of stage I oocytes is clearly visible but there is also a high level of signal in the remaining cytoplasm. During stage II, labeling is observed both in the cloud at the cortex and in the remaining cytoplasm. However, by stage III, no cytoplasmic labeling is detected, and all Xlerk RNA is localized to the vegetal pole. By stage IV, this labeling pattern is broader than that of either Xcat-2 or Xpat. C3H-3, also determined computationally to contain CAC-rich clusters in its 3â² UTR, utilizes the late, or Vg1, pathway. In this case, localization does not begin until stage II, and background labeling is observed through stage III. By stage IV of oogenesis, however, this late-pathway RNA shows no labeling in the animal hemisphere. Vg1 is shown as a known example of late-pathway localization, and histone mRNA (Hist), which is expressed throughout the cytoplasm of stage IâIV oocytes, is shown as a negative control for localization. The scale bar represents 100 μm in all panels.
Image published in: Betley JN et al. (2002)
Copyright © 2002. Image reproduced with permission of the Publisher, Elsevier B. V.
| Gene | Synonyms | Species | Stage(s) | Tissue |
|---|---|---|---|---|
| nanos1.L | nos1, xcat-2, xcat2, xnos1 | X. laevis | Sometime during oocyte stage I to oocyte stage IV | mitochondrial cloud vegetal cortex cytoplasm |
| pgat.L | Xpat | X. laevis | Sometime during oocyte stage I to oocyte stage IV | mitochondrial cloud vegetal cortex cytoplasm |
| efnb1.L | cfnd, cfns, efl3, elk-l, ephrinB1, eplg2, lerk2 | X. laevis | Sometime during oocyte stage I to oocyte stage IV | mitochondrial cloud vegetal cortex cytoplasm |
| zfp36l2.L | c3h-3, XC3H-3b, zfp36l2-a, zfp36l2-b | X. laevis | Sometime during oocyte stage I to oocyte stage IV | mitochondrial cloud vegetal cortex cytoplasm |
| gdf1.S | dvr-1, dvr1, gdf-1, vg-1, vg1 | X. laevis | Sometime during oocyte stage I to oocyte stage IV | mitochondrial cloud vegetal cortex cytoplasm |
Image source: Published
Permanent Image Page
Printer Friendly View
XB-IMG-44221