XB-IMG-121151
Xenbase Image ID: 121151
|
|
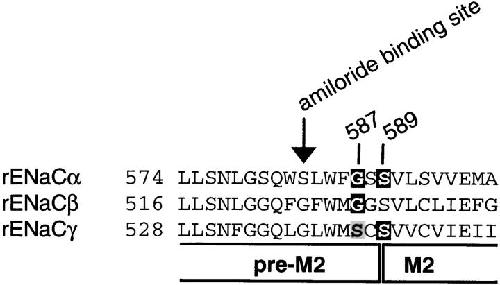
Figure 1. Sequence alignment of the preM2/M2 segments of rat α, β, and γ ENaC. The number of the first residue shown is indicated. The putative start of M2 and the position of the amiloride binding site (αS583, βG525, and γG537) are indicated. Mutations that change ionic selectivity are mutations of αG587 and αS589 as well as residues at the homologous positions in β and γ subunits. Residues shown in white on a dark background change Na+/K+ selectivity when mutated, and mutation of αG587 and the homologous residues in β and γ subunits change Li+/Na+ selectivity. Mutation of γS541 (shown in black on gray background) changes Li+/Na+, but not Na+/K+ selectivity. Image published in: Kellenberger S et al. (2001) © 2001 The Rockefeller University Press. Creative Commons Attribution-NonCommercial-ShareAlike license Larger Image Printer Friendly View |
